
Pingouin
Pingouin is an open-source Python package for statistics. It is built on top of Pandas and NumPy and provides high-level functions to perform the most common statistical tests. Learn more on the documentation or check the code on GitHub
Senior ML data scientist
Oura Ring
I am a neuroscientist specialized in sleep, health and wearables. I currently work as a Senior machine-learning data scientist at Oura ring, where I build the next generation of sleep algorithms.
Prior to this, I have worked for 5 years as a postdoctoral sleep researcher in the Center for Human Sleep Science at UC Berkeley (Prof. Matt Walker's lab).
My research examined how sleep — or the lack of it — impacts human health and diseases (including diabetes, atherosclerosis and Alzheimer's disease). My work has been published in top-tier journals and featured in several major news media.
My domains of expertise include machine learning & signal processing, particularly when applied to polysomnography and/or wearable physiological data (e.g. EEG, ECG, PPG, HRV).
I firmly believe in open science. I am the creator and core maintainer of several open-source libraries in Python,
which are used by thousands across the globe:
Pingouin (statistics, +500k downloads & taught in several universities),
YASA (automatic sleep staging) and
AntroPy (complexity of time-series).
In my free time, you can find me enjoying precious moments with my amazing wife and daughter, playing music with friends, cooking, or spending time outdoors.
| 2018-23 | Postdoctoral sleep researcher, Center for Human Sleep Science (Walker lab), University of California, Berkeley |
| 2014-17 | PhD in Neuroscience, with honors, Lyon 1 University, France | 2012-14 | Master degree in Neuroscience, cum laude, Lyon 1 University, France |
| 2009-12 | Bachelor degree of Cognitive Sciences, summa cum laude (ranked 1st), Lyon 2 University, France |
| Software development | Python, R, Matlab, SQL, cloud-computing (AWS), Git, Docker |
| Signal processing | Polysomnography (EEG, EKG), structural and functional MRI, accelerometer, photoplethysmography (PPG), heart rate variability (HRV) |
| Data science | Machine learning and deep learning, statistical modeling, data processing and visualization, time-series analysis |
| Machine-learning in Python, graduate level (UC Berkeley) |
| Mentoring of several undergraduate and postgraduate students (UC Berkeley) |
| Neurobiology, undergraduate level (Lyon University) |
| Ethics in medicine, medical school (Lyon University) |
| Neuro-imaging (fMRI), graduate level (Lyon University) |
Please find below a list of selected publications (last updated June 2022)
PDF versions are provided for individual, noncommercial purposes only. These files may not be reposted without permission. Copyright and all rights therein resides with the respective copyright holders, as stated
within each paper.

Please visit my GitHub repository for an exhaustive list of the projects/softwares that I am contributing to.

Pingouin is an open-source Python package for statistics. It is built on top of Pandas and NumPy and provides high-level functions to perform the most common statistical tests. Learn more on the documentation or check the code on GitHub

YASA (Yet Another Spindle Algorithm) is an open-source Python package for sleep analysis. It implements several state-of-the-art and validated algorithms to score sleep stages, detect spindles & slow-waves, and calculate features from sleep polysomnography recordings. Learn more on the documentation or check the code on GitHub
AntroPy is an open-source Python package for computing several entropy & complexity metrics of time-series. Learn more on the documentation or check the code on GitHub.
Please find below some tutorials regarding the analyses I specialize in.

Why do some people remember more of their dreams?
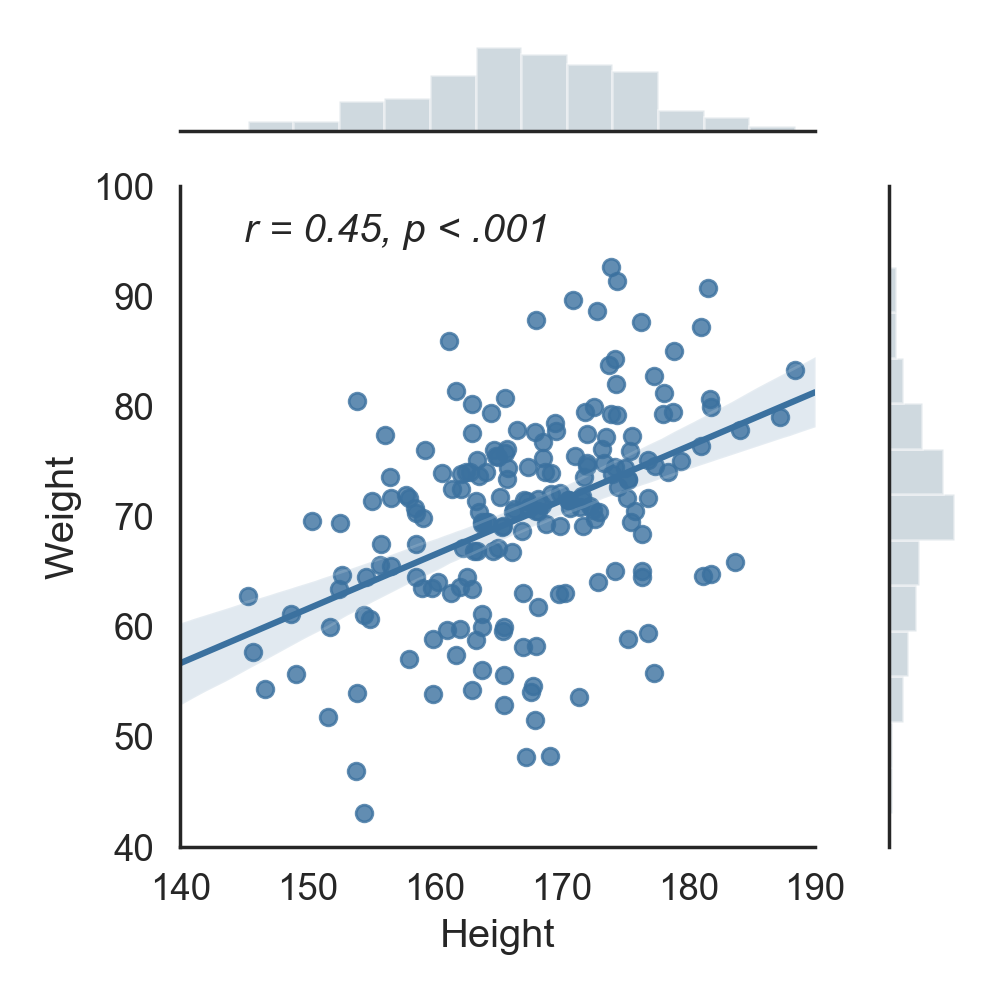
A primer on correlation and significance testing between two or more variables in Python.
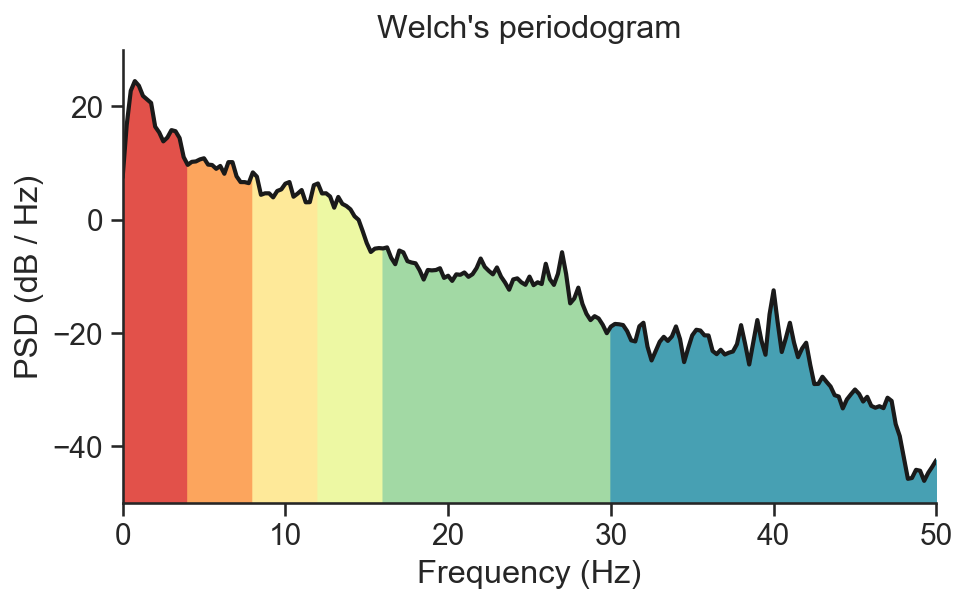
Compute the average bandpower of an EEG signal in Python.